1. 2022-2023: Prelude and origin from biology to reusable work
TAFFISH did not begin directly as a programming language or a Hub. Its 2022 materials are better understood as a prelude: the starting point was biological research, especially attempts to connect different omics layers in a meaningful analysis. During this work, a practical problem became visible: many steps consumed time and energy even though they were repetitive, transferable, and suitable for preservation.
That realization led to a more explicit tool idea in 2023. The first multi-omics tool plan, dated 2023-06-21, described multi-omics analysis as a field with many data types, scattered tools, inconsistent formats, and repeated manual integration work. The target was not yet a package manager or a DSL. It was a framework for preserving reusable procedures, bringing existing tools together, cleaning intermediate data, and supporting downstream biological interpretation.
A second milestone came with the 2023-08-27 three-step plan. The direction became more restrained: start from a single-omics system, especially Hi-C or 3D genome analysis, then generalize toward multi-omics workflows. That document already emphasized modularity: TAD and loop callers could be embedded, while the new work would focus on input, output, and intermediate format conversion.
This stage was still not TAFFISH, but several ideas that later survived were already present: tool integration, workflow reuse, modularity, reducing repeated work, and turning command-line procedures into reusable artifacts.
2. 2024: BioFlow and BioHub and the environment problem
The next obstacle was not whether a workflow could be written. It was whether the same workflow could survive another server, another operating system, or another software version. Around 2024 this pushed the project toward BioFlow and BioHub.
BioFlow focused on describing a single bioinformatics workflow. BioHub began to carry the meaning of a tool library and platform. The 2024 BioHub reports framed the core pain points clearly: installation was hard, command-line interfaces were inconsistent, tools were hard to combine, environments could conflict, and workflows were difficult to pass from one person to another.
This was the decisive turn. The project was no longer only about a multi-omics analysis method. It became a system for managing tools, workflows, environments, and standards.
3. 2024-10 to 2025-03: TAFFISH as a Shell-oriented DSL and Hub
From late 2024 to early 2025, BioHub and BioFlow evolved into TAFFISH. The name TAFFISH was settled in October 2024 during patent preparation, and a January 2025 report later made the name visible in the title “TAFFISH (2024)”. The project was then placed in the context of difficult installation, fragile environments, workflow construction, portability, reproducibility, collaboration, and inheritance.
TAFFISH was defined as Tools And Flows Framework Intensify SHell: a domain-specific language based on Shell for command-line workflows. By March 2025, the system was described as two coordinated parts: the TAFFISH language and the TAFFISH-HUB. The language described tools and flows. The Hub stored taf tools and taf flows so that users could install and run them in a manner reminiscent of apt, yum, conda, or pip.
The core syntax ideas were already visible: pass parameters with
command-line options, bind placeholders such as ::name::,
use tags such as <container:...> for runtime
environments, and expose apps as taf-xxx commands.
4. 2024-2025: Prototype, package manager, and early apps
By the 2024 summary and the early 2025 reports, TAFFISH had a working interpreter and a taf package manager. It supported Debian 12, Ubuntu 18.04.1 or newer, and Apple Silicon Macs. The early ecosystem already contained more than 20 taf apps, including bedtools, samtools, BLAST, bowtie2, fastp, juicer, STAR, subread, trim_galore, and example flows for RNA-seq and gene family search.
This proved the central claim: command-line bioinformatics tools could be wrapped as taf scripts, tied to containerized runtime environments, installed by a package manager, and invoked as normal commands.
It was still a prototype. Some early tags were too heavy, several features were broader than the core problem, and GUI experiments arrived before the CLI semantics were fully stable. Later design work deliberately reduced the system back toward a smaller center: keep Shell composability, add only the tags needed for environment and parameter semantics, and let the Hub grow from that stable core.
5. 2025: Ecosystem growth, GitHub migration, and preprint
In 2025, TAFFISH moved from a prototype into an expanding ecosystem. Reports from that year record the continued maintenance of the interpreter and package manager, broader operating-system support, and a growing app collection covering genomics, proteomics, transcriptomics, 3D genomics, base images, and selected GUI tools.
The Hub also changed shape. What began as a private deployment gradually moved toward GitHub repositories and static indexes. This made the system easier to publish, inspect, mirror, and reuse without maintaining a dedicated backend for every stage of the project.
The preprint TAFFISH: A lightweight, modular, and containerized workflow framework for reproducible bioinformatics analyses marked the project as a coherent research output rather than only a local engineering tool.
6. 2026: Common Lisp, LispWorks, and a full-system refactor
TAFFISH has been built in Common Lisp from its early stage. A Rust rewrite was considered for distribution and systems engineering, but the project returned to Common Lisp because the language fits TAFFISH's core work: DSL design, interpreter structure, compiler passes, interactive development, and rapid refactoring.
The latest version also introduced LispWorks as the Linux delivery route. This solved a major portability problem on Linux and removed the need to maintain multiple Linux-specific installation packages as in the old version. macOS packaging has not yet been carried into this new route, but remains a later adaptation target.
The 2026 refactor was not a narrow cleanup. It was a full-system
rebuild that reached from the compiler core to the app project
model, taf-cli, package metadata, Hub indexes, and the GitHub-based
publishing structure. The codebase was separated into clearer
layers: taffish-core for lexing, parsing, parameter
binding, emitters, and compilation; taffish-cli for
the terminal entry point; taf-core and
taf-cli for project management, installation, build,
and publish flows; and taffish-hub for the app and
flow ecosystem.
With Linux portability separated from app logic, the project could focus more directly on the language, package manager, app structure, and Hub index rather than on parallel installer maintenance.
A key change was the relationship between taf apps and Shell.
Earlier recursive taf-app calls could create compile-time and
runtime conflicts. The new model uses shell wrappers and delayed
compilation, so commands such as taf-test can
participate in ordinary pipelines:
echo 123 | taf-test cat
The <taffish> tag and [[taf: ...]]
syntax remain meaningful because they let a flow explicitly declare
TAFFISH app nodes. Their purpose is not merely to run commands, but
to compile related taf apps before a long flow starts, catching
missing apps, parameter problems, or compilation errors earlier.
7. 2026-05: Current positioning
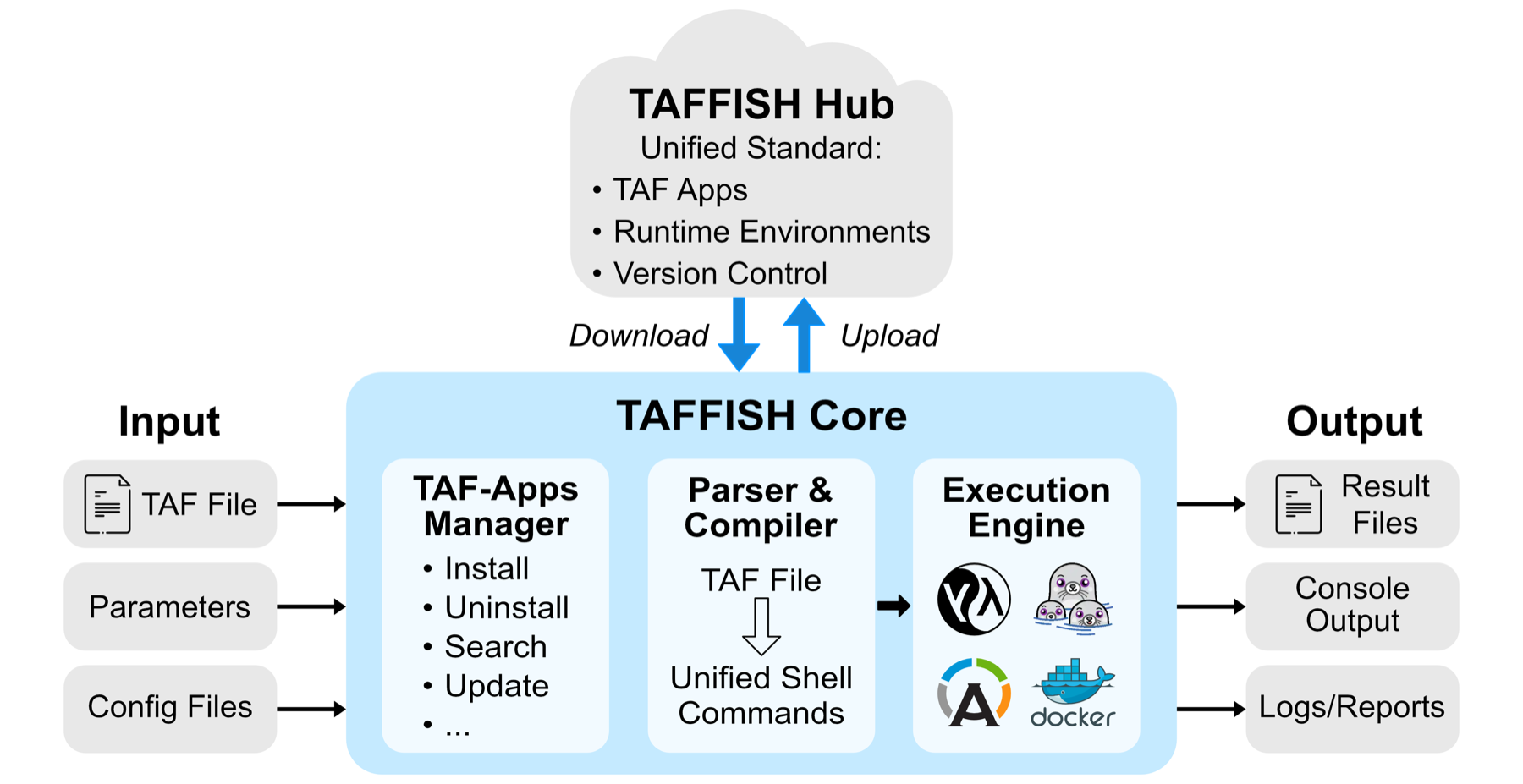
TAFFISH is now best understood as a reproducible command-line ecosystem for bioinformatics tools and workflows. Shell keeps the barrier to entry low and preserves composition. Containers bound the runtime environment. taf scripts describe tools and flows. The taf package manager installs and distributes apps. TAFFISH-HUB makes those apps discoverable through a static index.
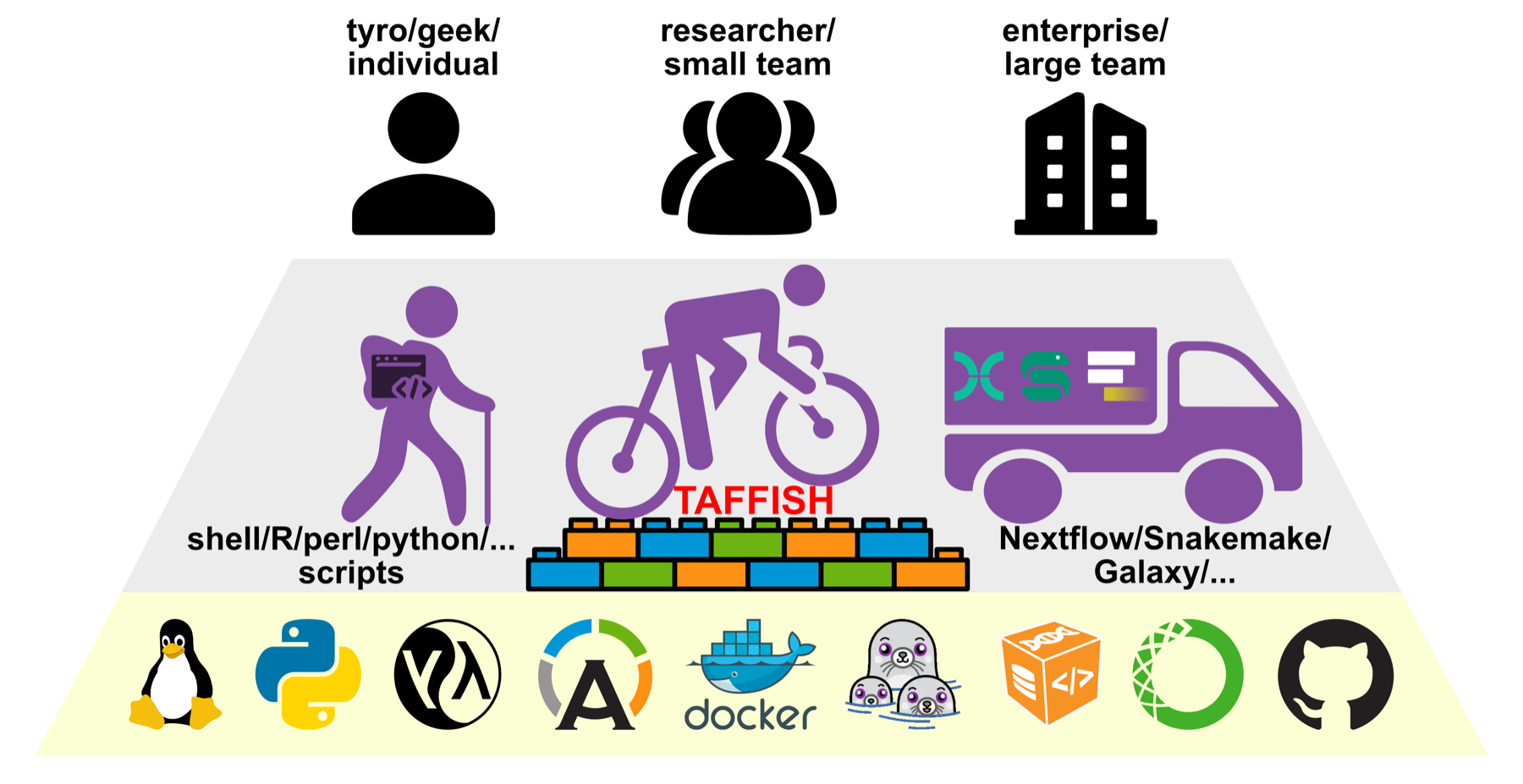
TAFFISH is not simply a replacement for Nextflow, Snakemake, or Galaxy, and it is not only a container launcher. It focuses on a more basic problem: making command-line tools exist in a more unified, reproducible, and inheritable form across systems, users, and projects.
This page was compiled on 2026-05-08. For future updates, keep the dated sections as historical records, add new milestones before this current-positioning section, and then rewrite this final section to reflect the latest stable view.